Diagnosis of Thyroid Nodules from Medical Ultrasound Images with Deep Learning
By Eunjung Lee, School of Mathematics and Computing (CSE), Yonsei University
Small lumps or growths on the thyroid gland are usually benign and cause no symptoms. A small percentage of these thyroid nodules, however, are malignant. Physicians use high-resolution ultrasonography1 to diagnose thyroid nodules, following up with a biopsy for nodules that exhibit common signs of malignancy; these include solidity, irregular margins, microcalcifications, and a shape that is taller than it is wide (Figure 1).
While the characteristics of malignant nodules are well established, diagnosing malignancy from ultrasound images remains a challenge. The accuracy of the diagnosis depends on the experience of the radiologist, and radiologists assessing the same nodule can arrive at different diagnoses.
Our research team at Yonsei University and Severance Hospital (Seoul, Korea) used MATLAB® to design and train convolutional neural networks (CNNs) to identify malignant and benign thyroid nodules. Diagnostic tests have shown that these CNNs perform as well as expert radiologists. We validated the CNNs against data sets from multiple hospitals, packaged them with a user interface, and deployed them as a web application. The application is used by medical students as part of their training and by experienced radiologists who need an objective second opinion on diagnoses.
Previous Machine Learning and Deep Learning Approaches
Before exploring the use of deep learning for thyroid nodule diagnosis, we tried conventional machine learning models. We performed feature engineering and applied a variety of machine learning methods available in MATLAB, including support vector machine (SVM) and random forest classification. The models performed about as well as a radiologist with 10 to 15 years of experience. Our goal was to develop software that would perform as well as experienced radiologists but provide consistent and objective results all the time, and so we began evaluating deep learning approaches.
One difficulty in using deep learning for medical imaging classification is the lack of available data. Medical data, including images, is protected by numerous privacy regulations, making it difficult to assemble a data set large enough to train a CNN. To address this challenge, we worked with 17 pretrained networks in MATLAB, including AlexNet, SqueezeNet, ResNet, and Inception.
We saw that each pretrained CNN classified an image in a slightly different way, and quickly realized that a combination would perform better than any single network. Combining networks proved difficult, however, because we had initially used a variety of programming languages and environments to create the networks. By using MATLAB, we would then have a single environment to preprocess images; design, train, and combine CNNs; analyze and visualize results; and deploy the CNNs as a web app.
Designing, Training, and Validating the CNNs
Before we began training CNNs in MATLAB, we performed several preprocessing steps on ultrasound images from four different hospitals in Korea. For example, we performed normalization to ensure that the pixel values of all the grayscale images were within the 0 to 255 range (Figure 2). For each image, we extracted a region of interest that focused on the nodule. We performed left-right mirroring on several hundred images of benign nodules to give us an equal number of benign and malignant nodules in our training data set.
We trained 17 different pretrained networks on a data set of more than 14,000 images that were approved by the institutional review boards (IRBs) of Severance Hospital: 4-2019-0163. Based on the performance of each, we chose a subset that included AlexNet, GoogLeNet, SqueezeNet, and InceptionResNetV2 to use in classification ensembles.
We experimented with two approaches for creating the ensembles, one that combined features and another that combined probabilities. For the feature-based combination, we used the outputs of the final fully connected layer in each CNN as features to train an SVM or random forest classifier. For the probability-based combination, we calculated a weighted average of the classification probability produced by each CNN. For example, if one CNN classified a nodule as benign with a 55% probability and another classified the same nodule as malignant with a 90% probability, then depending on the weighting, the ensemble was likely to classify the nodule as malignant.
To validate the diagnostic performance of the ensembles, we generated receiver operating characteristic (ROC) curves and compared the area under the curve (AUC) for each ensemble against the AUC for expert radiologists. We performed this comparison on the internal data set from Severance Hospital (part of the Yonsei University Health System) and on external data from three other hospitals. On the internal test set, the AUC of the AlexNet-GoogLeNet-SqueezeNet-InceptionResNetv2 ensemble was significantly higher than that of the radiologists. On the external test sets, the AUC was about the same (Figure 3).
Deploying the SERA Web App
To make our CNNs available in the hospitals that Yonsei University works with, we created a web app named SERA and deployed it with MATLAB Web App Server™. Accessible via a web browser, the SERA app is used for academic purposes only at present. Doctors use SERA to get a second opinion in the diagnostic process. The app is also used to train first- and second-year doctors.
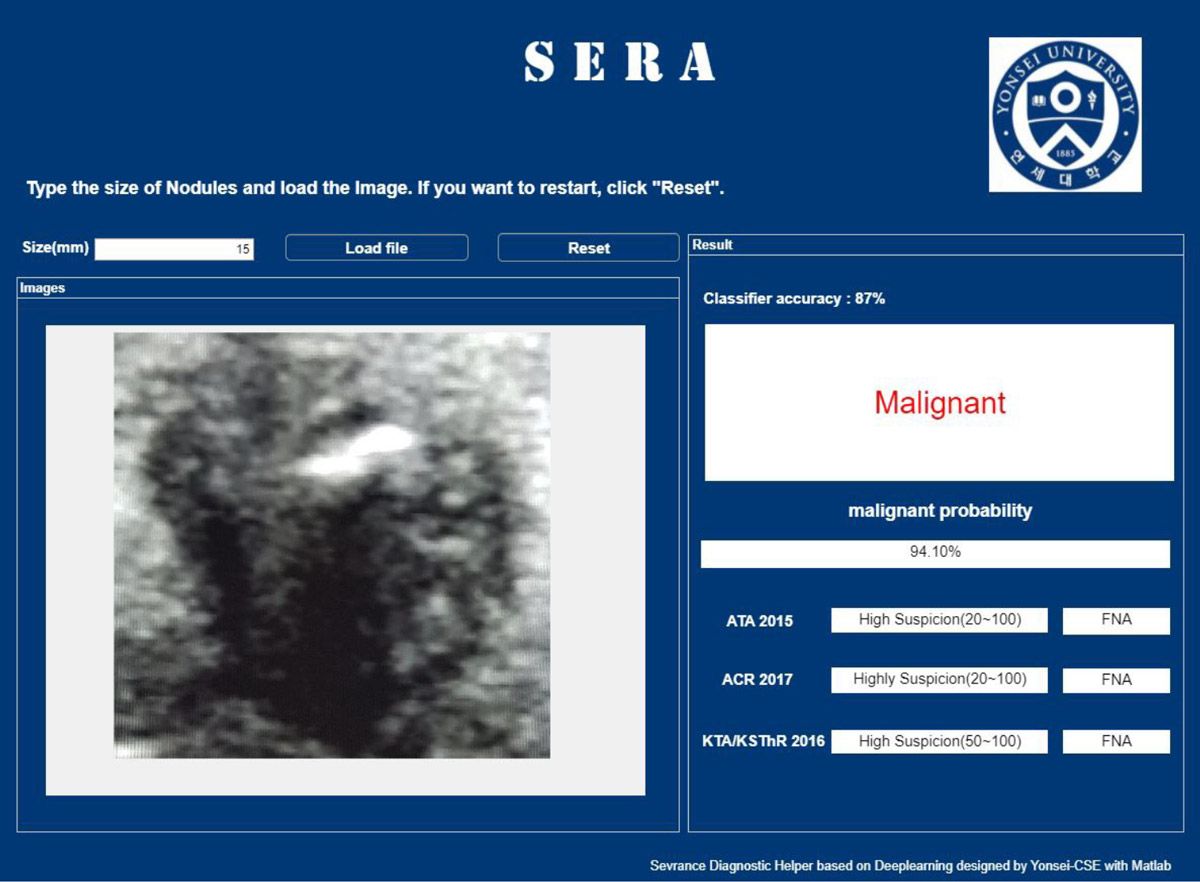
A simple user interface enables radiologists to run SERA on newly captured ultrasound images (Figure 4). Once an image is loaded, the CNNs diagnose the thyroid nodule in the image and the app displays a classifier accuracy score along with the probability that the nodule is malignant. Based on this probability, the app may recommend a fine needle aspiration (FNA) biopsy to confirm the diagnosis.
CNN Explainability and Extensibility
Our team is currently working on the explainability of our CNNs—that is, why and how they come to a decision when classifying thyroid nodules. To do this, we are studying specific layers in the CNNs—in particular, filters in the convolution layer—to understand which image features the network is using to make its decision. We plan to meet with our most experienced radiologists to determine to what degree highly trained humans and trained CNNs use the same image features in thyroid nodule diagnoses.
We are also planning to develop CNN-based applications for diagnosing breast cancer and skin cancer.
1 Ultrasonography images usually have low resolution. However, due to their noninvasive and less harmful properties compared with other medical imaging tools, high-resolution ultrasonography is widely used even for pregnant women and infants.
2 Koh, Jieun, Eunjung Lee, Kyunghwa Han, Eun‑Kyung Kim, Eun Ju Son, Yu‑Mee Sohn, Mirinae Seo, Mi‑ri Kwon, Jung Hyun Yoon, Jin Hwa Lee, Young Mi Park, Sungwon Kim, Jung Hee Shin, and Jin Young Kwak. “Diagnosis of thyroid nodules on ultrasonography by a deep convolutional neural network.” Scientific Reports 10, no. 1 (September 2020): 15245.
Published 2022