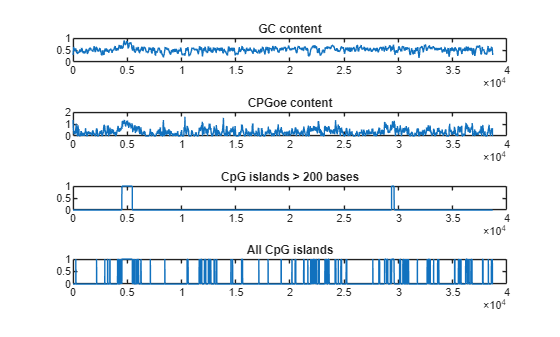
cpgisland
Locate CpG islands in DNA sequence
Description
cpgStruct = cpgisland(SeqDNA)cpgStruct, a MATLAB® structure containing the starting and ending bases of the CpG islands greater
than the minimum island size of 200 bases.
cpgStruct = cpgisland(SeqDNA,Name=Value)
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Version History
Introduced before R2006a