pcsegdist
Segment point cloud into clusters based on Euclidean distance
Syntax
Description
labels = pcsegdist(ptCloud,minDistance)minDistance between points from different clusters.
pcsegdist assigns an integer cluster label to each point in
the point cloud, and returns the labels of all points.
[
also returns the number of clusters.labels,numClusters] = pcsegdist(ptCloud,minDistance)
[___] = pcsegdist(___,
sets properties using name-value arguments. For example, Name=Value)labels =
pcsegdist(
sets the minimum and maximum number of points in each cluster to
ptCloud,minDistance,NumClusterPoints=[1,Inf])[1,Inf].
Examples
Create two concentric spheres and combine them.
[X,Y,Z] = sphere(100);
loc1 = [X(:),Y(:),Z(:)];
loc2 = 2*loc1;
ptCloud = pointCloud([loc1;loc2]);
pcshow(ptCloud)
title('Point Cloud')
Set the minimum Euclidean distance between clusters.
minDistance = 0.5;
Segment the point cloud.
[labels,numClusters] = pcsegdist(ptCloud,minDistance);
Plot the labeled results. The points are grouped into two clusters.
pcshow(ptCloud.Location,labels)
colormap(hsv(numClusters))
title('Point Cloud Clusters')
Load an organized lidar point cloud to the workspace.
ld = load('drivingLidarPoints.mat');Detect the ground plane. Distance is measured in meters.
maxDistance = 0.9; referenceVector = [0 0 1]; [~,inliers,outliers] = pcfitplane(ld.ptCloud,maxDistance,referenceVector);
Remove the ground plane points.
ptCloudWithoutGround = select(ld.ptCloud,outliers);
Cluster the point cloud with a minimum of 10 points per cluster using the exhaustive method.
minDistance = 2; minPoints = 10; [labels,numClusters] = pcsegdist(ptCloudWithoutGround,minDistance,Method="exhaustive",... NumClusterPoints=minPoints);
Remove the points with a label value of 0.
idxValidPoints = find(labels); labelColorIndex = labels(idxValidPoints); segmentedPtCloud = select(ptCloudWithoutGround,idxValidPoints);
Plot the labeled results.
figure
colormap(hsv(numClusters))
pcshow(segmentedPtCloud.Location,labelColorIndex)
title('Point Cloud Clusters')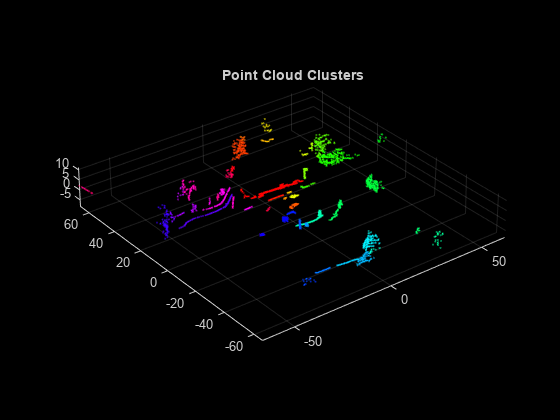
Input Arguments
Point cloud, specified as a pointCloud object.
Minimum Euclidean distance between points from two different clusters, specified as a positive scalar.
Data Types: single | double
Name-Value Arguments
Example: ParallelNeighborSearch=false sets
the ParallelNeighborSearch to
false.
Specify optional pairs
of arguments as Name1=Value1,...,NameN=ValueN, where
Name is the argument name and Value is the
corresponding value. Name-value arguments must appear after other arguments, but the
order of the pairs does not matter.
Minimum and maximum number of points in each cluster, specified as a
scalar or a 2-element vector of the form
[minPoints,maxPoints]. When
you specify NumClusterPoints as a scalar, the
maximum number of points in the cluster is unrestricted. The function
sets labels to 0 when clusters
are outside of the specified range.
Method to segment the point cloud, specified as
"approximate" or "exhaustive".
Set Method to "exhaustive" to
ensure that no points outside of each cluster are less than
minDistance away. This approach uses the
density-based spatial clustering of applications with noise (DBSCAN)
algorithm. Set Method to
"approximate" for faster segmentation, but at the
expense of accuracy.
Parallel neighbor search to segment point cloud data, specified as
true or false. Set this
property to true when you expect there to be
approximately 50 clusters or more with fewer than 100 points per
cluster.
A parallel neighbor search can improve segmentation speed for some
datasets. Improved speed depends on the dataset and the value of the
minDistance input. This argument is not
supported when you set the Method name-value
argument to "exhaustive".
Output Arguments
Cluster labels, returned as one of the following.
If the point cloud,
ptCloud, stores point locations as an unorganized M-by-3 matrix, thenlabelsis an M-by-1 vector.If the point cloud,
ptCloud, stores point locations as an organized M-by-N-by-3 matrix, thenlabelsis an M-by-N matrix.
Each point in the point cloud has a cluster label, specified
by the corresponding element in labels. The value of
each label is an integer from 0 to the number of clusters
of valid points, numClusters. The value
0 is reserved for invalid points, such as points with
Inf or NaN coordinates.
Data Types: uint32
Number of clusters, returned as a positive integer. The number of clusters
excludes the label value 0, which is reserved for invalid
points. The function returns numClusters as a
single data type when the value of the
Location property of the ptCloud
object is single. Otherwise, the function returns the
value as a double data type.
Data Types: single | double
Extended Capabilities
The "exhaustive" segmentation method, which can be set by using
the Method name-value argument, does not support C/C++ code
generation.
Usage notes and limitations:
The generated CUDA® code segments the point cloud into clusters by using a combination of algorithms described in [1] and [2]. The output from the generated code can differ slightly with results from MATLAB® simulation.
The
"exhaustive"segmentation method, which can be set by using theMethodname-value argument, does not support GPU code generation.
References
[1] Andrade, Guilherme, Gabriel Ramos, Daniel Madeira, Rafael Sachetto, Renato Ferreira, and Leonardo Rocha. “G-DBSCAN: A GPU Accelerated Algorithm for Density-Based Clustering.” Procedia Computer Science 18 (2013): 369–78. https://doi.org/10.1016/j.procs.2013.05.200.
[2] Kalentev, Oleksandr, Abha Rai, Stefan Kemnitz, and Ralf Schneider. “Connected Component Labeling on a 2D Grid Using CUDA.” Journal of Parallel and Distributed Computing 71, no. 4 (April 2011): 615–20. https://doi.org/10.1016/j.jpdc.2010.10.012.
Version History
Introduced in R2018aStarting in R2026a, pcsegdist function supports the
NumClusterPoints name-value argument for GPU code
generation.
Configure the Method name-value argument to choose between
"exhaustive" or "approximate"
segmentation methods. The ParallelNeighborhood name-value
argument does not support the "exhaustive" method.
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
Seleccione un país/idioma
Seleccione un país/idioma para obtener contenido traducido, si está disponible, y ver eventos y ofertas de productos y servicios locales. Según su ubicación geográfica, recomendamos que seleccione: .
También puede seleccionar uno de estos países/idiomas:
Cómo obtener el mejor rendimiento
Seleccione China (en idioma chino o inglés) para obtener el mejor rendimiento. Los sitios web de otros países no están optimizados para ser accedidos desde su ubicación geográfica.
América
- América Latina (Español)
- Canada (English)
- United States (English)
Europa
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)