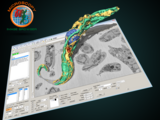
Microscopy Image Browser (MIB)
MIB is a package for segmentation of multi-dimensional (2D-4D) microscopy datasets
herehttp://se.mathworks.com/matlabcentral/fileexchange/63402-microscopy-image-browser-2--mib2-With MIB you can analyse, segment and visualize various multidimensional datasets from both light and electron microscopy. See more further details and tutorials on MIB website
- 1,3K (desde el inicio)
- 7 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
9 feb 2019
Microscopy Image Browser 2 (MIB2)
MIB2 is an update package for segmentation of multi-dimensional (2D-4D) microscopy datasets
With MIB2 you can analyse, segment and visualize various multidimensional datasets from both light and electron microscopy. MIB2 is completely rewritten to follow MVC architecture and brings
- 2,4K (desde el inicio)
- 6 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
26 sep 2025
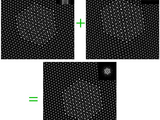
High-Throughput, Algorithmic Determination of Nanoparticle Structure from Electron Microscopy Images
Automated analysis of electron microscopy images (PC and Mac versions available.)
. Chem. Soc. 2014, 136, 7603 doi: 10.1021/ja503509kHigh-Throughput, Algorithmic Determination of Nanoparticle Structure from Electron Microscopy ImagesChristine R. Laramy, Keith A. Brown, Matthew N
- 840 (desde el inicio)
- 2 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
25 abr 2017
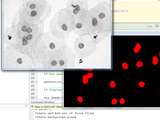
Segmentation of WBC in microscopy images
Stain deconvolution followed by threshold based segmentation is used.
- 207 (desde el inicio)
- 1 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
21 nov 2021
Read .dm3 and .dm4 image files
Read Digital Micrograph files for electron microscopy
- 3K (desde el inicio)
- 13 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
20 sep 2013
Filter noise and interpolate microscopy images in frequency domain
Remove spatial frequencies beyond the optical cutoff and perform physically accurate interpolation.
. Filtering the spatial frequencies beyond the optical cutoff provides simple yet effective means of reducing noise. Since microscopy data is band-limited, padding in frequency domain provides accurate
- 881 (desde el inicio)
- 1 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
1 mar 2013
Imagic, MRC, DM and STAR file i/o
Functions for reading and writing image files and STAR files for electron microscopy
This directory contains m-functions for reading and writing files used in electron microscopy and 3D reconstruction. The file formats those used by the IMAGIC software package (Image Science GmbH
- 3,9K (desde el inicio)
- 10 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
6 ene 2019
Atomic Force Microscopy Image Analysis
This is the software to analysis the atomic force microscopy images. Average feature size and area can be calculated using this software. Steps to follow - 1 - First remove any additional part of the
- 2,6K (desde el inicio)
- 2 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
7 dic 2010
GUI for displaying image stacks, e.g. time-resolved or z-stacked microscopy images.
- 1,8K (desde el inicio)
- 2 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
1 sep 2011
Image to Volume Conversion for Representing Materials Topography
Converts a single image (microscopy, topogrpahy) into array of images.
- 3,8K (desde el inicio)
- 1 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
17 nov 2003
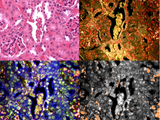
Machine learning assisted hyperspectral imaging
Automatic detection of nanoparticles using hyperspectral microscopy and machine learning
Automatic detection of nanoparticles using hyperspectral microscopyNanoparticles are used extensively as biomedical imaging probes and potential therapeutic agents. As new particles are developed and
- 720 (desde el inicio)
- 2 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
20 abr 2017
Computationally Enhanced Quantitative Phase Microscopy
Toolbox for the ceQPM method
. "Computationally Enhanced Quantitative Phase Microscopy Reveals Autonomous Oscillations in Mammalian Cell Growth." bioRxiv (2019): 631119.The image processing pipeline includes:1. load the experiment information2
- 74 (desde el inicio)
- 1 (últimos 30 días)
- -- / 5
- Comunidad
-
6 may 2020
Transmittance limit for Digital Holographic Microscopy
Estimates and plots the transmittance limit for a digital hologram in a 1-bit to 16-bit representation for the given setup parameters.
Retrieval of accurate phase maps from Digital Holographic Microscopy (DHM) is highly dependent on the representation of the carrier fringe pattern in the digital sensor. Physically, the contrast of
- 19 (desde el inicio)
- 1 (últimos 30 días)
- -- / 5
- Comunidad
-
24 ene 2024
Noise evaluation in transmission electron microscopy (TEM)
Scrips for evalutating the important noise parameters and correlations in TEM. Reconstruction of the point spread function from signal noise
microscopy.Sci Rep 15, 3815 (2025).https://doi.org/10.1038/s41598-025-85982-4So, if you use any of these scripts for your evaluations please cite the above paper.Please note, however, that the authors disclaim any
- 11 (desde el inicio)
- 1 (últimos 30 días)
- -- / 5
- Comunidad
-
16 jun 2025
- 308 (desde el inicio)
- 4 (últimos 30 días)
- -- / 5
- Comunidad
-
11 sep 2024
This is a tutorial demonstrating how to access open microscopy data and publish reusable and reproducible analyses in MATLAB
Analyse Open Microscopy Data in MATLAB®A MATLAB Live Script with accompanyingJupyter® Notebook,m fileandreproducible code capsule on Code Ocean®to access and analyze Microscopy image data sets from
- 21 (desde el inicio)
- 1 (últimos 30 días)
- -- / 5
- Comunidad
-
14 mar 2024
Synthetic Bright-Field Microscopy Image Generator
This is a synthetic image generation tool that can create realistic Pap-smear images
This is a simulator able to procedurally create realistic bright-field microscopy images depicting Pap-smears. The principles used in the simulation are described in the paper "Simulation of
- 618 (desde el inicio)
- 1 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
12 mar 2015
Particle localisation using local gradients
Local_gradientsThis package provides a set of tools for 3-D localisation of single particles in brightfield and fluorescent microscopy using local gradients.The package is provided in LabVIEW, Matlab
- 35 (desde el inicio)
- 2 (últimos 30 días)
- -- / 5
- Comunidad
-
9 nov 2022
A Bayesian Adaptive Basis Algorithm for Single Particle Reconstruction
3D reconstruction algorithm for electron cryo-microscopy.
a Bayesian maximum-a-posteriori framework and uses an efficient optimization algorithm for the maximization. Evaluations using simulated and actual cryogenic electron microscopy data show resolution
- 939 (desde el inicio)
- 1 (últimos 30 días)
- -- / 5
- Comunidad
-
5 abr 2012
Nonlinear Drift Correction from Scanning Probe Microscopy (SPM) Orthogonal Image Pairs
Correct nonlinear drift distortions in scanning probe images.
This collection of scripts is intended to correct nonlinear drift distortions in images recorded using any scanning probe microscopy technique (where there is a slow and a fast scan direction). It
- 562 (desde el inicio)
- 1 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
26 feb 2016
SubspaceEM: A Fast Maximum-a-posteriori Algorithm for Cryo-EM Single Particle Reconstruction
Fast alignment and reconstruction algorithm for cryo-electron microscopy
cryo-electron microscopy images and the structure projections, greatly reducing the number of image transformations and comparisons that are computed. The files include an implementation of the SubspaceEM
- 415 (desde el inicio)
- 1 (últimos 30 días)
- -- / 5
- Comunidad
-
18 mar 2015
Segment the vessel branches from dynamic image of fluorescent microscopy
Segment the blood vessels from a dynamic image of fluorescent microscopy. == Install ======- Add all attached files to matlab path- Download "Better Skeletonization" from following URL and add to
- 2,3K (desde el inicio)
- 3 (últimos 30 días)
- -- / 5
- Comunidad
-
5 abr 2012
Allows users to view a tiff stack (useful for time-lapse microscopy datasets)
- 494 (desde el inicio)
- 2 (últimos 30 días)
- -- / 5
- Comunidad
-
19 feb 2015
MATLAB code for a Graphic User Interface dedicated to automatically count and quantify dendritic spines from fluorescence microscopy images
MATLAB code for a Graphic User Interface dedicated to automatically count and quantify dendritic spines from fluorescence microscopy images. It has been tested with .oib files from
- 58 (desde el inicio)
- 2 (últimos 30 días)
- -- / 5
- Comunidad
-
18 nov 2020
Optical Point Spread Function generator (with aberrations)
3D PSF of a microscopy objective starting Focal length, stop size and numerical aperture. Includes Aberrations, and illumination properties.
- 81 (desde el inicio)
- 2 (últimos 30 días)
- -- / 5
- Comunidad
-
19 sep 2024
Stain Deconvolution is used to determine Stain density Estimation in H&E microscopy images
- 73 (desde el inicio)
- 1 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
21 nov 2021
Hyperspectral CARS Image Analysis
Hyperspectral CARS microscopy and spectroscopy toolbox
Hyperspectral CARS microscopy and spectroscopy toolbox allows researchers easy analysis of their data.Toolbox focuses on image fusion, denoising and spectroscopy.
- 743 (desde el inicio)
- 1 (últimos 30 días)
- 2,0 / 5
- Comunidad
-
23 sep 2014
Measure Myelin-to-Axon Diameter
The M-file is to calculate the g-ratio for Scanning or Transmission Electronic Microscopy.
- 277 (desde el inicio)
- 1 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
6 ene 2017
Segmentation of neurons in microscopy images
Set up, train and apply a neural network for segmentation of neurons in microscopy images.
- 64 (desde el inicio)
- 1 (últimos 30 días)
- -- / 5
- Comunidad
-
13 ago 2021
GUI for detection of cells for Integrated Light and Electron Microscopy
GUI for detection of fluorescent cells in In-Resin Fluorescence sections
Matlab GUI for automated detection of fluorescent cells in In-Resin Fluorescence sections for Integrated Light and Electron Microscopy
- 101 (desde el inicio)
- 1 (últimos 30 días)
- -- / 5
- Comunidad
-
12 feb 2018
Composites fibre orientation from one cross section
Computes the 2nd order orientation tensor diagonal from an optical or scanning electron micrograph
- 287 (desde el inicio)
- 1 (últimos 30 días)
- 2,5 / 5
- Comunidad
-
27 jun 2016
Automated Analysis of Platelet Aggregation in Microfluidics
Automatic thresholding of fluorescence microscopy images in microfluidics to determine platelet coverage and aggregate size distribution.
- 42 (desde el inicio)
- 1 (últimos 30 días)
- -- / 5
- Comunidad
-
18 nov 2019
Interface detection in point cloud data (SRM & SMLM)
Code for detecting interfaces in super-resolution microscopy (SRM) data. Also works for other types of Poisson-distributed point cloud data.
SMLM_interface_detectionCode for detecting interfaces in super-resolution microscopy (SRM) data,typically single-molecule localization microscopy (SMLM).The code provided here is under Copyright ©
- 6 (desde el inicio)
- 1 (últimos 30 días)
- -- / 5
- Comunidad
-
10 abr 2022
AxonSeg is a GUI that performs axon and myelin segmentation on histology images.
Segment axon and myelin from microscopy data. Written in Matlab. The compiled versions are also available for those who do not have the necessary processing toolboxes.
- 1,7K (desde el inicio)
- 2 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
12 jun 2019
Automatic spatial drift correction of images or video frames.
each image frame aligned. An instruction manual has been submitted to Microscopy Today and will hopefully be published in the near future.
- 907 (desde el inicio)
- 1 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
10 feb 2014
Extract and display ROIs from a Zeiss CZI image
Two programs that extract and display ROIs from Zeiss CZI images.
- 145 (desde el inicio)
- 3 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
24 mar 2021
STXM Spectromicroscopy Particle Analysis Routines
Spectromicroscopic analysis of atmospheric nanoparticles (aerosols)
An automated data analysis method for atmospheric particles using scanning transmission X-ray microscopy coupled with near edge X-ray fine structure spectroscopy (STXM/NEXAFS). This method is applied
- 1,3K (desde el inicio)
- 1 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
19 oct 2010
Leica image format file loader
Read and preview of Leica's image format file.
- 1,1K (desde el inicio)
- 2 (últimos 30 días)
- 4,7 / 5
- Comunidad
-
4 abr 2012
Finds, summarizes, and plots (if desired) 2-color colocalization from 3D images.
This script allows the user to load 3-D TIFF images, such as those derived from confocal or 2-photon microscopy, into the MATLAB workspace for analysis of colocalization between two images in 3-D
- 2,7K (desde el inicio)
- 2 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
18 ago 2004
Fast, complete two-photon pipeline
- 766 (desde el inicio)
- 9 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
18 ago 2016
Best Infocus Image from Image Stack
This code Find out the best infocus image from a image stack using Tamura coefficient.
This matlab code implements Tamura Coefficient to find out the best infocus image in the stack of the images.The stack of the microscopy has many images but one of the image is the best
- 66 (desde el inicio)
- 2 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
26 dic 2019
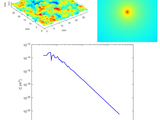
Radially averaged surface roughness/topography power spectrum (PSD)
Calculates radially averaged 2D power spectrum for a surface roughness/topography
surface topography, which the topography is normally obtained by any 3D profilometry techniques, such as AFM (Atomic Force Microscopy), WLI (White Light Interferometry) and many other optical profilers. As
- 3,4K (desde el inicio)
- 13 (últimos 30 días)
- 4,9 / 5
- Comunidad
-
19 dic 2016
Save Matlab data from SPM/AFM scans for use with Gwyddion
What is Gwyddion?"Gwyddion is a modular program for SPM (scanning probe microscopy) data visualization and analysis. Primarily it is intended for analysis of height fields obtained by scanning probe
- 586 (desde el inicio)
- 4 (últimos 30 días)
- 4,8 / 5
- Comunidad
-
22 abr 2014
Basic (demo) 2-photon microscope scanning software
- 194 (desde el inicio)
- 1 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
16 abr 2024
Read ROIs and ROI sets saved from ImageJ into MATLAB, without java.
- 4,5K (desde el inicio)
- 24 (últimos 30 días)
- 4,9 / 5
- Comunidad
-
16 ago 2017
3D-Reconstruction algorithm used for N-way FRET microscopy
- 182 (desde el inicio)
- 1 (últimos 30 días)
- 1,0 / 5
- Comunidad
-
28 dic 2015
STXM data analysis script collection - STACKLab
STXM data analysis script collection with stack exploration GUI tool STACKLab
This is a MATLAB script collection developed at Lawrence Berkeley National Lab that can be used as a basis for Scanning Transmission X-ray Microscopy (STXM) data analysis. It includes routines for
- 1,8K (desde el inicio)
- 1 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
13 jul 2009
Control ZEN Blue and the microscope from MATLAB
Control the ZEN Blue microscope control software from within a MATLAB script
- 357 (desde el inicio)
- 2 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
19 sep 2016
DM3 Import for Gatan Digital Micrograph
Imports Gatan .DM3 format files with tags (images, spectra, & spectral images) into a MATLAB struct.
This script acts to import files from Gatan's .DM3 file format, utilized for electron microscopy, into a MATLAB structure. The fields of the MATLAB structure can then be referenced with the
- 2,7K (desde el inicio)
- 4 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
8 ene 2013
Novel Retinal Vessel Segmentation Algorithm: Fundus Images
The algorithm presented here segments retinal blood vessels with a high degree of accuracy.
of Fungal Hyphae in Macroscopic Microscopy Image Stacks." arXiv preprint arXiv:1704.02356 (2017).Saranya, M., and A. Grace Selvarani. "Fundus Image Screening for Diabetic Retinopathy." Indian Journal
- 8,7K (desde el inicio)
- 9 (últimos 30 días)
- 4,6 / 5
- Comunidad
-
25 mar 2019
Script package for quantifying bacterial load within cells using images from an ArrayScan microscope
This package includes a variety of scripts for analysis of microscopy images, using the SAFIRE screening platform. This package is useful for analyzing high-content screens of intracellular bacteria
- 46 (desde el inicio)
- 1 (últimos 30 días)
- -- / 5
- Comunidad
-
20 feb 2017
Reads and imports Asylum Research ARDF files to Matlab structures for force curve analysis.
For atomic force microscopy force curve analysis within Matlab. Reads Asylum Research Data Files (.ARDF).readARDF() reads image, note, and other data from the ARDF file into a Matlab
- 242 (desde el inicio)
- 3 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
15 mar 2024
Nikon ND2 File Reader
- 896 (desde el inicio)
- 5 (últimos 30 días)
- 4,5 / 5
- Comunidad
-
25 abr 2019
Medical Imaging Toolbox Interface for Cellpose Library
Use Cellpose library in MATLAB
Segment cells from microscopy images using the Medical Imaging Toolbox™ Interface for Cellpose Library. The support package provides functionality for downloading pretrained models from the Cellpose
- 482 (desde el inicio)
- 12 (últimos 30 días)
- -- / 5
- MathWorks
-
26 ene 2026
Current version only accept gray-scale images. Just format your image stack into a 3D array.
A good tool to display all kinds of 3D image stacksLSM (Laser scanning microscopy) imagesCT scan (x-ray) imagesMRI imagesConfocal microscopy imagesOCT (optical coherence tomography) images
- 1,4K (desde el inicio)
- 1 (últimos 30 días)
- -- / 5
- Comunidad
-
25 oct 2012
Time-lapse microscopy classification of 2D cells based on cell shape (Seiler et al., TERM, 2012)
- 1,3K (desde el inicio)
- 1 (últimos 30 días)
- 1,0 / 5
- Comunidad
-
16 jul 2012
Read and write files in SPIDER format
SPIDER is a free image processing system for electron microscopy. It is used for three-dimensional reconstruction of single particle macromolecules, multivariate statistical classification, and
- 903 (desde el inicio)
- 1 (últimos 30 días)
- 4,0 / 5
- Comunidad
-
19 may 2009
Diffusion gradient vector field in 3D
Generate in 3D the diffusion gradient vector field as in Xu and Prince 1998
usage:[un vn wn] = dgvf_calc(I,sqrt(numel(I)), 0.5, 1, 1, 1, 1)Note: The code was developed with 3d flourescence microscopy images in mind (bright objects against dark background). Enforcement of boundary
- 920 (desde el inicio)
- 1 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
6 ene 2014
Radially averaged surface roughness power spectrum (PSD) only on top or bottom part of a topography
Calculates radially averaged 2D power spectrum for a certain part of surface topography
topography is normally obtained by any 3D profilometry techniques, such as AFM (Atomic Force Microscopy), WLI (White Light Interferometry) and many other optical profilers.You need to provide 5 inputs to the
- 1K (desde el inicio)
- 5 (últimos 30 días)
- 5,0 / 5
- Comunidad
-
19 dic 2016
DiffractIndex is a simple program for measuring diffraction ring diameters from TEM or XRD patterns
- 932 (desde el inicio)
- 2 (últimos 30 días)
- 4,2 / 5
- Comunidad
-
7 ene 2016