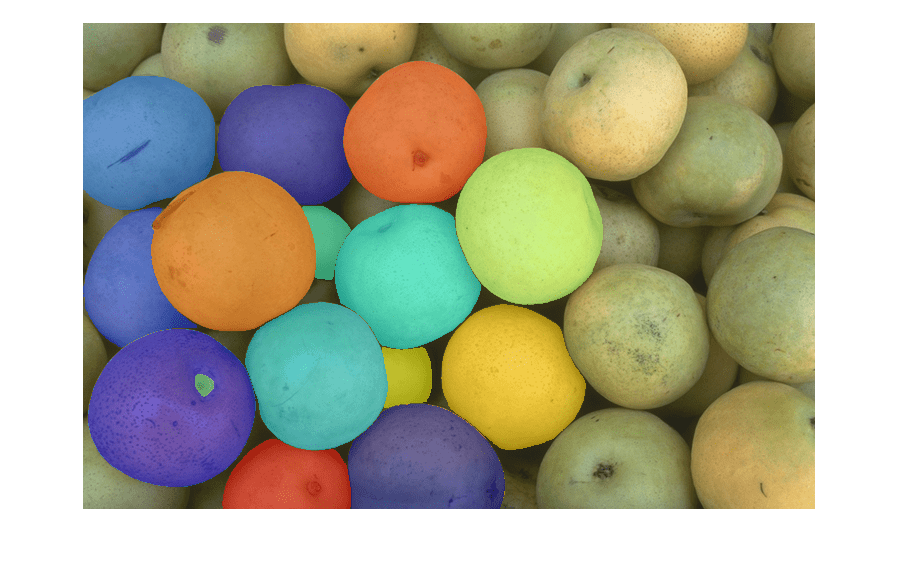imsegsam
Perform automatic full image segmentation using Segment Anything Model 2 (SAM 2)
Since R2024b
Description
Add-On Required: This feature requires one of these add-ons.
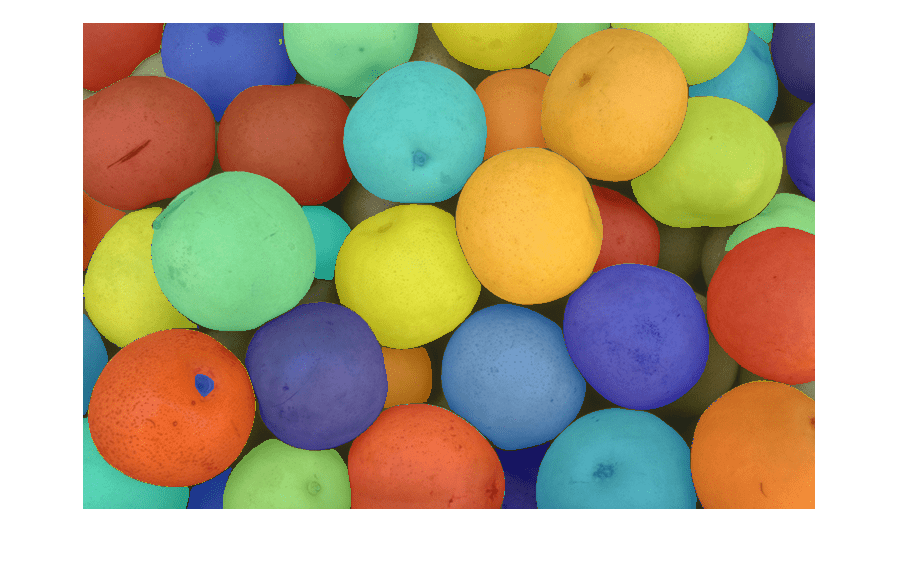
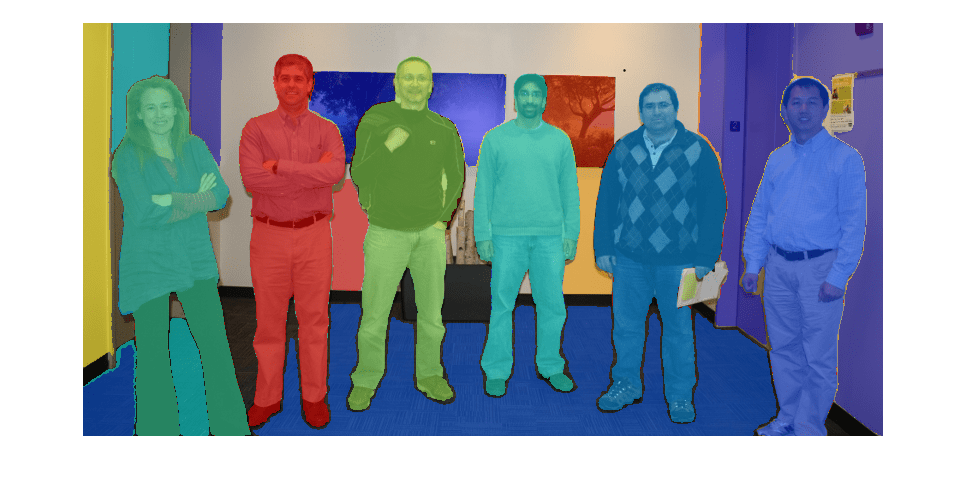
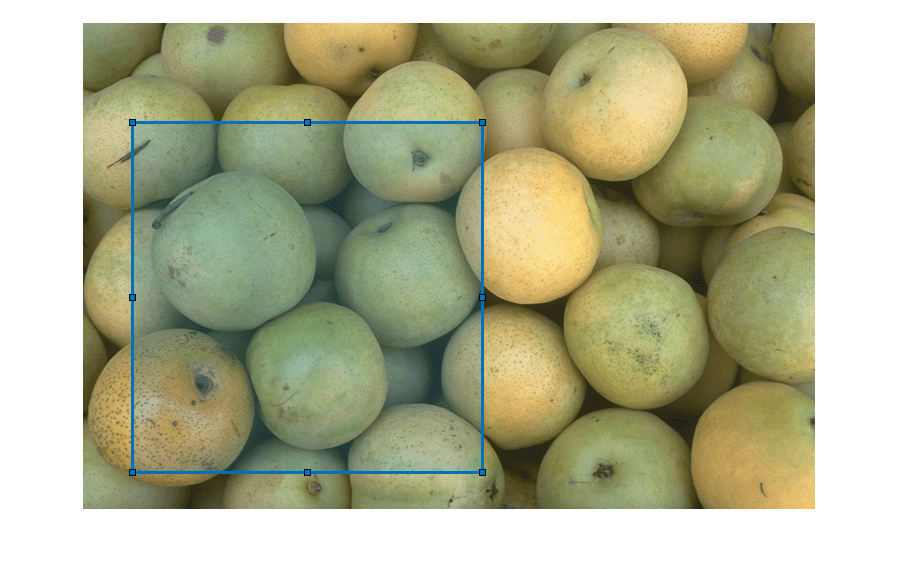
Use the imsegsam function to automatically segment an entire
image or all of the objects inside a region of interest (ROI) using the Segment Anything Model
2 (SAM 2) or Segment Anything Model (SAM). The function samples a regular grid of points on an
image and returns a set of predicted masks for each point, which enables the model to produce
multiple masks for each object and its subregions. You can customize various segmentation
settings based on your application, such as the ROI in which to segment objects, the size
range of objects which to segment, and the confidence score threshold with which to filter
mask predictions.
Note
To use any of the SAM 2 models, this functionality requires the Image Processing Toolbox™ Model for Segment Anything Model 2 add-on if you use any of the SAM 2 models. To use the base SAM model, this functionality requires the Image Processing Toolbox Model for Segment Anything Model add-on.
[
specifies options using one or more name-value arguments. For example,
masks,scores] = imsegsam(I,Name=Value)PointGridSize=[64 64] specifies the number of grid points that the
imsegsam function samples along the x- and
y- directions of the input image as 64
each.
Examples
Input Arguments
Name-Value Arguments
Output Arguments
Tips
For best model performance, use an image with a data range of [0, 255], such as one with a
uint8data type. If your input image has a larger data range, rescale your image to the range [0, 1] by using therescalefunction and then convert the image to theuint8data type by using theim2uint8function.To visualize object masks, you can display the masks as a label matrix or a stack of binary masks.
To create a label matrix from the
masksstructure, use thelabelmatrixfunction. For an example of how to visualize masks using a label matrix, see the Customize Full Image Segmentation example.To create a stack of binary masks, convert the
masksstructure to a logical array. For an example of how to create and display a stack of masks, see the Segment Objects as Mask Stack Using Segment Anything Model.
References
[1] Kirillov, Alexander, Eric Mintun, Nikhila Ravi, Hanzi Mao, Chloe Rolland, Laura Gustafson, Tete Xiao, et al. "Segment Anything," April 5, 2023. https://doi.org/10.48550/arXiv.2304.02643.
[2] Ravi, Nikhila, Valentin Gabeur, Yuan-Ting Hu, Ronghang Hu, Chaitanya Ryali, Tengyu Ma, Haitham Khedr, et al. “SAM 2: Segment Anything in Images and Videos.” arXiv, October 28, 2024. https://doi.org/10.48550/arXiv.2408.00714.
Version History
Introduced in R2024bSee Also
Functions
segmentAnythingModel|labelmatrix|labeloverlay|insertObjectMask(Computer Vision Toolbox)
Apps
- Image Segmenter | Image Labeler (Computer Vision Toolbox)